Highlighted Publication
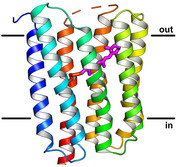
Cryo-EM structure of the bacterial light-driven proton pump proteorhodopsin
Please see our publication: Hirschi et al. (2021), Nature Commun.
Highlighted Publication
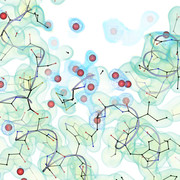
Crystal structure of the L-arginine/agmatine transporter AdiC at 1.7 Å resolution: Insights into the role of water molecules and networks in oligomerization and substrate binding
Please see our recent publication: Ilgü et al. (2021), BMC Biology
Welcome to the website of the Fotiadis Lab
Structure and function of membrane proteins
We investigate the structure and function of membrane proteins with a special focus on membrane transport proteins. The methods of choice are high-resolution microscopy techniques (cryo-electron and atomic force microscopy) and X-ray crystallography.
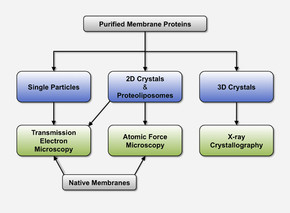
Methods
Our methods of choice for the structure determination of membrane proteins are electron microscopy and X-ray crystallography. In addition atomic force microscopy is used to image membrane proteins embedded in lipid bilayers under near-physiological conditions.
Our mission
Our mission is to assess the structure and function of membrane proteins in the detergent-solubilized state and in their native environment, the lipid bilayer.
Target proteins
Our target membrane proteins consist mainly of membrane transport proteins and receptors.
Funding
Swiss National Science Foundation (SNSF) Project funding, National Centre in Competence in Research (NCCR) TransCure and NCCR Molecular Systems Engineering, and Sinergia.
Latest Tweets from FotiadisLab @IBMM_UniBE
March 7, 2024:
Dear all, please check out our pretty cool story in Advanced Science about the
light color-controlled pH-adjustment of aqueous solutions using engineered proteoliposomes.
Congrats to Daniel Harder and our collaborators in Basel, Noah Ritzmann and Daniel J. Müller.
https://onlinelibrary.wiley.com/doi/full/10.1002/advs.202307524
February 2, 2023:
Finally out!
Please have a look at our cryo-EM structure of the canine distemper virus (CDV) attachment glycoprotein (H protein). Amazing architecture of this morbillivirus ectodomain.
https://doi.org/10.1073/pnas.2208866120
February 24, 2022:
Peptide transporter structure reveals binding and action mechanism of a potent PEPT1 and PEPT2 inhibitor.
Please check out our newest publication by Stauffer and colleagues:
https://www.nature.com/articles/s42004-022-00636-0
Congratulations Mirko, Jean-Marc, Hüseyin, Zöhre and Rajendra !
November 12, 2021:
Excellent collaboration between the University of Bern, the Zurich University of Applied Sciences (ZHAW), the University of Lyon/INSERM and the EPFL.
Please check it out:
https://journals.asm.org/doi/10.1128/mBio.02621-21
October 21, 2021:
Interested in L-lactate transport inhibitors ?
Then please check out our recent publication by Patrick Bosshart and colleagues in Communications Chemistry:
https://www.nature.com/articles/s42004-021-00564-5?proof=t%29
Congratulations Patrick !
October 6, 2021:
For membrane protein aficionados:
Want to apply for an Eccellenza Professorial Fellowship from the Swiss National Science Foundation and need a host institute ?
Then check out the possibility at the Institute of Biochemistry and Molecular Medicine:
https://ohws.prospective.ch/public/v1/jobs/31373766-098e-4e3c-bda5-2c08bc7bf855
September 9, 2021:
Our crystal structure of the L-arginine/agmatine transporter AdiC at 1.7 Å resolution is out !
The high-resolution structure gives new insights on the role of water in oligomerization and substrate binding.
A big thank you to the NCCR TransCure !
September 3, 2021:
Please check out our latest publication within the NCCR Molecular Systems Engineering:
Engineering and functional characterization of a proton-driven β-lactam antibiotic translocation module for bionanotechnological applications
Congratulations Mirko!
July 7, 2021:
Please check out our latest publication:
Cryo-EM structure and dynamics of the green-light absorbing proteorhodopsin
Congratulations to all authors and a special "thank you" to Stephan (UniBE) and Thomas (UniZH/ETHZ) !!!
https://www.nature.com/articles/s41467-021-24429-6
June 16, 2021:
Open Postdoc position in the FotiadisLab (Bern, Switzerland):
In structural characterization of viral and membrane proteins by cryo-electron microscopy
For details, please see:
https://ohws.prospective.ch/public/v1/jobs/57f4dcac-73d4-4f15-baaa-87250ca47353
January 27, 2021:
Open positions at the FotiadisLab
Ph.D. student position:
https://ohws.prospective.ch/public/v1/jobs/bdf4bcc1-bb86-4397-acda-85f5b75b6c0d
Posdoc position:
https://ohws.prospective.ch/public/v1/jobs/87f12943-1613-4fc2-bb61-2a4fc6f815f9
Are you interested in membrane proteins biochemistry, function and structure ?
We are looking for researchers to join our team !
October 19, 2020:
Please check out our newest publication on the effects of the ancillary protein 4F2hc on the substrate affinity and specificity of the human transporters LAT1 and LAT2.
Great work by Satish and colleagues, congratulations !
https://www.mdpi.com/1422-0067/21/20/7573
October 12, 2020:
Our first contribution to Coronavirus research.
An excellent collaboration with Thomas Lemmin (ETH Zurich/University of Zurich) and Philippe Plattet (Vetsuisse, University of Bern) – thank you !
https://www.sciencedirect.com/science/article/pii/S2590152420300209?via%3Dihub
October 1, 2020:
One step further...
Please check out Jean-Marc’s new publication entitled: “Sub-Nanometer Cryo-EM Density Map of the Human Heterodimeric Amino Acid Transporter 4F2hc-LAT2”.
Excellent job !
https://www.mdpi.com/1422-0067/21/19/7094
September 29, 2020:
Please discover our new double-chambered device for macromolecular crystal flash-cooling in different cryogenic liquids by Jeckelmann et al.
Congratulations Jean-Marc !
https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0239706
August 26, 2020:
Great review by Patrick Bosshart (@bossi1983) and colleagues (@MouseTransLab, @AvedaSci, @SchlessingerLab, @IBMM_UniBE) on SLC16 family transporters. Congrats !
https://www.sciencedirect.com/science/article/pii/S096800042030178X
August 19, 2020:
Very happy that we: Philippe Plattet (Uni Bern), Markus Seeger (Uni Zurich) and I (Uni Bern) received one of the 28 new Coronavirus grants funded by the National Research Programme COVID-19.
Thanks to this funding, we will be able to contribute in the fight against Coronavirus.
August 13, 2020:
Want to produce engineered versions of 2D bacteriorhodopsin crystals (purple membranes) for basic research and applied technologies ?
Then, please check out our recent publication in Methods Protoc.: https://pubmed.ncbi.nlm.nih.gov/32707904/
Thank you Mirko and colleagues for the great job!
March 26, 2020:
Check out our latest publication within the NCCR Molecular Systems Engineering:
Cryo-electron microscopic and X-ray crystallographic analysis of the light-driven proton pump proteorhodopsin reveals a pentameric assembly
https://www.sciencedirect.com/science/article/pii/S2590152420300064
Congratulations Stephan !
March 4, 2020:
Check out our latest publication:
Cryo-EM structure of the prefusion state of canine distemper virus fusion protein ectodomain
https://www.sciencedirect.com/science/article/pii/S2590152420300039
Congratulations David !
June 14, 2019:
First structure of an SLC16 solute carrier family transporter with bound L-lactate and inhibitor: www.nature.com/articles/s41467-019-10566-6
Thank you and congratulations Patrick, David and Sara - you did a great job !
Our research is founded by